Drug-induced liver injury is the leading cause of acute liver failure in the United States and a major driver of drug withdrawals from the market and failures in late-stage clinical development. It is also, by a significant margin, one of the hardest drug safety liabilities to predict preclinically.
That combination makes DILI worth understanding in depth — not just for toxicologists and safety pharmacologists but for anyone involved in drug development decisions. This post explains what DILI actually is, why it behaves so differently from most other drug toxicities, and why predicting it reliably remains one of the genuinely hard problems in preclinical safety science.
What DILI Is
Drug-induced liver injury is hepatic damage caused by a drug, its metabolites, or an immune response triggered by drug exposure. It covers a broad spectrum of presentations: from asymptomatic elevations in liver enzymes that resolve on drug withdrawal, to fulminant hepatic failure requiring liver transplantation.
Clinically, DILI is classified in several ways. The most important distinction for preclinical prediction is between intrinsic and idiosyncratic DILI.
Intrinsic DILI is dose-dependent and predictable. Given a high enough dose of acetaminophen, virtually everyone will develop liver injury. The mechanism is well understood: high-dose acetaminophen overwhelms hepatic glutathione stores, and the reactive metabolite NAPQI covalently binds to cellular proteins, causing oxidative stress and hepatocyte death. Intrinsic DILI is detectable in conventional preclinical models because it occurs at doses that can be tested experimentally and through mechanisms that are pharmacologically predictable.
Idiosyncratic DILI is the harder problem. It occurs in a small fraction of exposed patients, often at therapeutic doses, with no clear dose-response relationship, and frequently with a delayed onset ranging from days to months after drug initiation. It is not predictable from the pharmacology of the drug and is not reliably detected in animal models or conventional in vitro assays. Watkins (2011) described idiosyncratic DILI as "the black swan of hepatotoxicity" — rare, unpredictable, and capable of causing catastrophic outcomes in the individuals it affects.
Idiosyncratic DILI is responsible for most of the late-stage clinical failures and post-marketing drug withdrawals that make DILI such a costly problem for drug development. It is the form of DILI that preclinical science has historically been least equipped to predict.
Why Predicting DILI Is So Difficult
The dose-response problem
Most drug toxicities follow a dose-response relationship that preclinical models can, in principle, detect. DILI often does not. Idiosyncratic DILI can occur at or below therapeutic doses in susceptible individuals while the same drug is tolerated without any liver signal in the vast majority of patients. An animal study or cell-based assay run at a therapeutically relevant concentration may show no signal at all, while a small proportion of clinical patients will develop serious liver injury at the same exposure.
This is not a failure of the assay. It is a fundamental feature of idiosyncratic toxicity: the susceptibility is in the patient, not in the drug dose.
The mechanistic heterogeneity problem
DILI is not one thing mechanistically. It encompasses direct hepatocellular toxicity, cholestatic injury, steatosis, mixed hepatocellular-cholestatic patterns, and immune-mediated reactions that involve the drug or a drug-protein adduct as an antigen. Different mechanisms require different assay systems to detect. A viability assay will detect direct cytotoxicity. It will not detect cholestasis. A bile transport assay will detect drugs that impair BSEP function. It will not detect mitochondrial toxicity. A metabolic activation assay will detect reactive metabolite formation. It will not detect immune-mediated idiosyncrasy.
The DILIrank dataset maintained by the FDA classifies over 1,000 drugs by their DILI potential across multiple severity tiers. The mechanistic diversity within that dataset is striking: the same DILI severity classification can encompass drugs with completely different mechanisms of hepatic injury. Predicting DILI requires, in principle, covering all of these mechanisms simultaneously — which no single assay system can do.
The metabolism problem
Many hepatotoxic compounds are not toxic themselves but are converted to reactive or toxic species by hepatic metabolism. This metabolic activation means that the toxicity is mediated by a metabolite that may not be present at all in cell lines with low CYP expression, may be present at different levels depending on donor-specific CYP polymorphisms, and may be further processed by Phase II enzymes in ways that either detoxify it or generate secondary reactive species.
HepG2, the most widely used hepatocyte cell line, expresses CYP450 enzymes at roughly 1/1000th of the activity found in primary human hepatocytes. Compounds that cause DILI through metabolic activation will appear non-toxic in HepG2-based assays and reveal their liability only in more metabolically competent systems — such as pixHep™, Pixl Bio's iPSC-derived hepatocytes engineered for primary-like CYP activity and metabolic function, built on over a decade of hepatocyte differentiation expertise developed at DefiniGEN, the Cambridge-based iPSC pioneer whose merger with Phenaros formed Pixl Bio. Xu et al. (2008) estimated that metabolic activation accounts for a substantial fraction of hepatotoxic compounds that are not detected in standard in vitro screens.
The genetic susceptibility problem
The strongest risk factor for idiosyncratic DILI, in many cases, is not the drug or the dose but the patient. Specific HLA alleles have been robustly associated with idiosyncratic DILI for a number of drugs: HLA-B57:01 for flucloxacillin-associated liver injury, HLA-A33:01 for terbinafine, HLA-DRB1*15:01 for amoxicillin-clavulanate. Pirmohamed et al. (2015) reviewed the genetic basis of drug hypersensitivity reactions including liver injury, documenting the expanding catalogue of pharmacogenomic associations.
Beyond HLA, polymorphisms in drug-metabolising enzymes, transporters, and mitochondrial function genes all contribute to inter-individual differences in DILI susceptibility. A preclinical model that does not capture this genetic diversity will not capture the biological substrate of idiosyncratic susceptibility. This is one reason Pixl Bio's approach to DILI prediction runs compounds across multi-donor panels of pixHep™ hepatocytes — capturing the population-level variability that single-donor experiments systematically miss.
The immune component problem
For a subset of DILI, the mechanism is not direct toxicity but an adaptive immune response to a drug or drug-protein adduct. The drug acts as a hapten, forming covalent bonds with hepatic proteins, and the resulting adducts trigger an immune response in genetically susceptible individuals. This mechanism is largely invisible to cell-based assays that lack an immune component, and is poorly modelled in standard animal studies.
The danger hypothesis, elaborated in the context of DILI by Daly and Day (2012), suggests that reactive metabolites — by causing cell stress and damage-associated molecular pattern release — may provide the danger signal that licenses immune activation. This framework helps explain why some reactive metabolite-forming drugs cause idiosyncratic DILI while others do not: the difference may lie not just in metabolite reactivity but in the degree of cellular stress and danger signal generated.
What Current Preclinical Models Can and Cannot Do
Animal models
Rodent and non-human primate models have been the backbone of preclinical hepatotoxicity assessment for decades. They detect intrinsic, dose-dependent hepatotoxicity reasonably well. They are substantially worse at detecting idiosyncratic DILI.
The reasons are multiple. Species differences in CYP expression and substrate specificity mean that the metabolic activation profile of a drug in rodents may differ substantially from its profile in humans. Genetic diversity in the animal model is constrained by the use of inbred strains. The immune component of idiosyncratic DILI is not well modelled in standard rodent studies. And the rare nature of idiosyncratic DILI means that even a large animal study would have limited statistical power to detect it.
Olson et al. (2000) reported that preclinical studies in rodents and non-human primates detected only around 50–70% of hepatotoxins that caused human liver injury, with substantial false positive rates for human-relevant DILI assessment.
Conventional in vitro models
Standard hepatocyte viability assays and enzyme inhibition screens detect direct cytotoxicity and some forms of metabolic disruption, but miss the majority of clinically relevant DILI mechanisms. They are run at concentrations that may not reflect the hepatic exposure achieved in patients, often without accounting for protein binding, tissue distribution, or metabolic accumulation. They rarely incorporate immune components. And they almost never account for the inter-individual genetic variation that underlies susceptibility.
The predictive performance of conventional in vitro DILI assays has been extensively evaluated. Blomme and Will (2016) reviewed the state of in vitro DILI prediction and concluded that while progress had been made in detecting certain mechanisms, no single assay or assay combination had demonstrated sufficient sensitivity and specificity to serve as a reliable preclinical DILI screen. The field has since moved toward multi-modal platforms — which is precisely what Pixl Bio's pixCellServices is designed to deliver.
Where the Field Is Moving
The past decade has seen substantial investment in more sophisticated approaches to DILI prediction, motivated by the continued failure of conventional preclinical models to protect patients and drug development programmes from late-stage hepatotoxicity surprises.
Multi-parametric imaging and morphological profiling. Cell Painting and related high-content imaging approaches capture morphological changes that are invisible to viability assays. Sub-lethal changes in mitochondrial morphology, ER stress markers, lysosomal accumulation, and cytoskeletal organisation can all be detected morphologically at concentrations that cause no measurable cell death. These early-stage morphological signatures may provide earlier and more sensitive detection of hepatotoxic potential than endpoint viability measurements. Pixl Bio's pixCellPaint platform is built specifically around this principle — automated high-content imaging combined with AI-driven feature extraction to generate phenotypic fingerprints at scale.
Metabolically competent human hepatocyte systems. iPSC-derived hepatocytes with improved CYP expression, primary hepatocyte spheroids with extended functional windows, and liver-on-chip microphysiological systems all aim to address the metabolic competence gap in conventional in vitro models. Bhise et al. (2016) demonstrated that liver-on-chip systems could maintain primary hepatocyte function for extended periods and detect hepatotoxins that conventional 2D cultures missed.
Multi-donor population models. Using panels of iPSC-derived hepatocytes from genetically diverse donors allows the population-level distribution of hepatotoxic susceptibility to be explored in vitro. Compounds that produce a strong response in a subset of donors may be flagging an idiosyncratic liability that would be invisible in a single-donor experiment. This is a core design principle of the pixHep™ platform.
Bile transport and cholestasis assays. BSEP inhibition is a well-validated mechanism of drug-induced cholestasis, and dedicated transport assays have significantly improved detection of cholestatic DILI liabilities. Dawson et al. (2012) provided an important early characterisation of BSEP inhibition as a predictor of cholestatic DILI risk. Pixl Bio's PFIC2 disease model — iPSC-derived hepatocytes with bile transport and polarity defects — enables direct investigation of cholestatic mechanisms in a human-relevant system.
Machine learning on clinical and preclinical data. The DILIrank dataset and related curated databases have enabled the training of machine learning models that integrate multiple structural, physicochemical, and assay features to generate probabilistic DILI risk scores. Chen et al. (2016) demonstrated that ensemble machine learning models could significantly outperform individual assay-based predictions for a held-out set of DILI-positive and DILI-negative compounds. Pixl Bio's AI-driven phenomics layer applies the same principle to morphological data — combining pixCellPaint imaging with predictive modelling to surface DILI signals that functional assays alone would miss.
Why DILI Prediction Will Remain Difficult
It is worth being honest about the ceiling.
True idiosyncratic DILI, in its immune-mediated forms, may be fundamentally resistant to prediction by any in vitro system that lacks a fully functional adaptive immune response. The HLA associations that predict risk for specific drugs are population genetics signals that can inform patient selection but cannot be replicated in a cell-based assay.
The best realistic goal for preclinical DILI prediction is probably not the elimination of clinical DILI surprises but the earlier and more reliable detection of the mechanisms that account for the largest fraction of observed clinical cases. Metabolically activated intrinsic toxicity, mitochondrial liability, cholestatic mechanisms, and reactive metabolite formation can all, in principle, be detected by appropriately designed in vitro systems. The idiosyncratic immune-mediated tail of the DILI distribution remains a harder problem.
Progress will come from combining multiple assay modalities rather than searching for a single predictive system, from incorporating genetic diversity into in vitro testing, and from applying machine learning to integrate signals across assay platforms. It is precisely this multi-modal approach — iPSC-derived hepatocytes, phenomic profiling, and AI-driven analysis — that underpins Pixl Bio's DILI prediction platform.
It will not come quickly. But it will come.
Built on a Decade of iPSC Hepatocyte Science
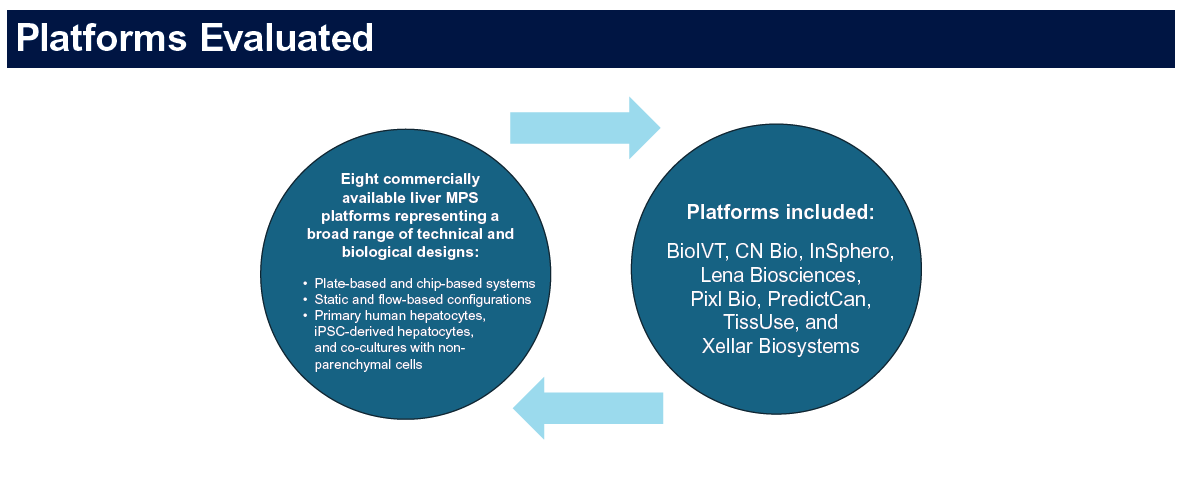
pixHep™ is the product of over ten years of iPSC differentiation science developed at DefiniGEN — a Cambridge-based company that pioneered the use of iPSC-derived hepatocytes for drug safety and disease modelling, and whose technology formed the biological foundation of Pixl Bio following the merger with Phenaros. DefiniGEN's hepatocyte models were deployed by pharmaceutical partners across DILI screening, metabolic liver disease modelling, and regulatory toxicology programmes — including as one of eight selected platforms in the FDA-CDER and 3Rs Collaborative cross-platform DILI study, one of the most rigorous multi-platform evaluations of liver microphysiological systems conducted to date, now accepted into the FDA's ISTAND qualification programme.
That depth of validation is what distinguishes pixHep™ from newer entrants to the iPSC hepatocyte space. The platform's functional breadth — primary-like CYP450 activity, albumin secretion, bile transport, and clinically anchored biomarker readouts including ALT and AST — was not established in a single study but refined across years of pharmaceutical partnerships and independent scientific scrutiny. Disease variants including MASLD, A1ATD, and PFIC2 extend the platform into the chronic and genetic liver disease space, where the same metabolic and functional rigour applies.
That scientific heritage is now Pixl Bio's — integrated with Phenaros's high-throughput phenomics and AI infrastructure to deliver a DILI prediction platform that combines human-relevant biology with the scale and analytical depth that modern drug development requires.
Pixl Bio develops predictive DILI models using iPSC-derived hepatocytes, Cell Painting phenomics, and AI analytics. To discuss how our approach could strengthen your safety programme, book a call with our team.
Tags: DILI · Drug-Induced Liver Injury · Hepatotoxicity · Drug Safety · Preclinical Safety · iPSC Hepatocytes · Cell Painting · Toxicology · CYP450 · Idiosyncratic Toxicity