This guide was developed by the pixlbio team. The Cell Painting methodology and AI phenomics platform at pixlbio were built by our scientific co-founders Jordi Carreras-Puigvert and Ola Spjuth, who together bring over two decades of research at the intersection of image-based phenomics, machine learning, and drug discovery. Jordi is Associate Professor at Uppsala University and co-author of the 2025 Nature Methods Cell Painting review. Ola is Professor of Pharmaceutical Bioinformatics at Uppsala University, creator of CPSign, and has 5,600+ citations across AI, conformal prediction, and large-scale Cell Painting research.
Cell Painting has quietly become one of the most powerful unbiased profiling technologies in modern drug discovery. Yet many biologists encounter it for the first time at a conference, read a paper referencing morphological profiling, and struggle to find a single clear resource that explains what it actually is, how it works, and how to apply it to their research.
This guide is our attempt to fix that.
Between us, we have spent over two decades working at the intersection of image-based phenomics, machine learning, and drug discovery — from fundamental methodology development to large-scale industrial applications at pixlbio. Our research group at Uppsala University helped establish Cell Painting as a national screening service at SciLifeLab, has published extensively on mechanism of action prediction, AI-driven phenomics, and conformal prediction models for drug safety, and continues to push the boundaries of what's biologically interpretable from high-content imaging data.
This guide draws on the work of the broader Cell Painting community — particularly the Carpenter-Singh lab at the Broad Institute, the JUMP-CP consortium, and the many researchers who have contributed open tools, datasets, and protocols. We have tried to represent their contributions accurately and encourage you to explore their work directly via the links throughout.
What is Cell Painting?
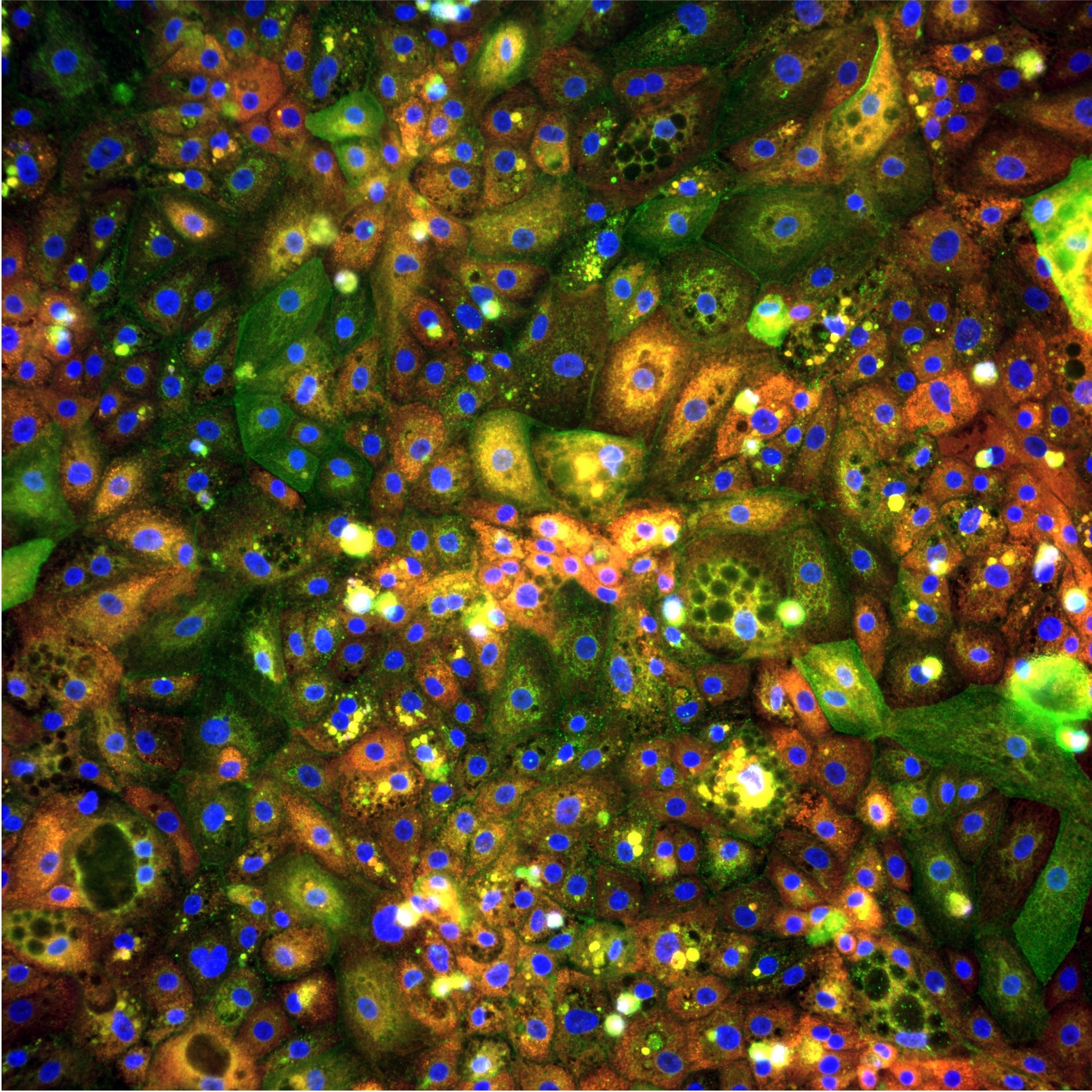
Cell Painting is a high-content, multiplexed fluorescence imaging assay designed to capture the morphological state of cells across multiple biological compartments simultaneously. It was first introduced by Gustafsdottir et al. (2013) in PLOS ONE and standardised into the protocol used across the field today by Bray et al. (2016) in Nature Protocols — one of the most cited papers in phenomics. Cell Painting has since become a cornerstone technology for image-based profiling in academic and industrial drug discovery.
The name comes from the concept of "painting" cells with a carefully chosen panel of fluorescent dyes, each targeting a distinct cellular structure, to create a rich multi-dimensional portrait of cell state.
Unlike targeted assays that measure one or two specific markers, Cell Painting captures thousands of morphological features per cell in a single experiment. This makes it uniquely powerful for unbiased phenotypic profiling — you don't need to know in advance what biology is important. The data tells you.
The Standard Dye Panel: What Gets Stained
The original Cell Painting panel uses six fluorescent dyes across five imaging channels, each targeting a distinct cellular compartment:
Together these dyes capture the state of virtually every major cellular compartment — nucleus, cytoplasm, organelles, membranes, and cytoskeleton. The resulting morphological fingerprint reflects the integrated biological state of the cell.
The standard six-dye panel is not fixed. Researchers regularly modify it to target different biology — adding markers for apoptosis, lipid accumulation, specific organelle states, or cell-type identity markers. The principle of unbiased multiplexed morphological profiling applies regardless of the specific dyes used.
How Cell Painting Works: Step by Step
Step 1 — Experimental design and cell treatment
Cells are seeded into microplates — typically 96-well or 384-well — and exposed to perturbations of interest:
- Small molecule compounds — for drug screening, mechanism of action studies, toxicity profiling
- Genetic perturbations — siRNA knockdowns, CRISPR knockout or activation screens
- Biological stimuli — cytokines, pathogens, metabolic stressors, disease-relevant conditions
- Combination perturbations — drug + genetic, compound synergy studies
The choice of cell model is flexible. Cell Painting has been applied to cancer cell lines, primary cells, stem cell-derived models, co-cultures, organoids, and patient-derived samples. At pixlbio, we have developed and validated Cell Painting workflows specifically for our iPSC-derived hepatocyte models (pixHep) — integrating human-relevant cell biology with phenomics at scale. The combination of a primary-like cell model with high-content imaging is, in our view, where the most meaningful drug discovery data comes from.
Step 2 — Fixation and staining
Cells are fixed, most commonly with 4% paraformaldehyde, and stained with the fluorescent dye panel. Dye concentrations, incubation times, temperature, and washing conditions all influence staining quality. Automation of the staining procedure is strongly recommended — manual staining introduces variability that is difficult to remove computationally downstream.
Step 3 — High-content fluorescence imaging
Fixed and stained plates are imaged using an automated fluorescence microscope. Multiple fields of view per well are captured — typically 9–16 fields at 20x magnification. A single 384-well plate generates thousands of individual images. Common imaging platforms include the PerkinElmer Opera Phenix, Molecular Devices ImageXpress, and Yokogawa CellVoyager series.
Step 4 — Image quality control and pre-processing
Raw images require pre-processing before analysis: illumination correction, background subtraction, focus quality filtering, and well-level QC flagging wells with low cell count, cell death, or staining failure. This step is often underestimated but is critical — poor QC propagates errors through all downstream analysis.
Step 5 — Cell segmentation
Individual cells are identified from images through segmentation — determining the boundaries of each cell for per-cell feature extraction. Approaches range from classical algorithms to modern deep learning:
- CellProfiler — the open-source standard with robust classical segmentation pipelines
- Cellpose — widely adopted deep learning segmentation model
- StarDist — deep learning segmentation optimised for star-convex shapes
- CellSAM — segment anything model adapted for biological cells
Step 6 — Morphological feature extraction
For each segmented cell, hundreds to thousands of numerical features are extracted quantifying intensity, texture, shape, radial distribution, neighbour relationships, and organelle co-localisation. The standard tool is CellProfiler, developed at the Broad Institute. A typical pipeline extracts 1,000–2,000 features per cell across all channels.
Deep learning-based feature extraction — using convolutional neural networks or vision transformers — is increasingly used as an alternative. Foundation models pretrained on large Cell Painting datasets can capture complex morphological patterns difficult to define manually.
Step 7 — Profile aggregation and normalisation
Per-cell features are aggregated to well level, then normalised to remove systematic variation — DMSO normalisation, robust Z-scoring, and spherize/whitening. Batch effects are the most common technical challenge at scale. The pycytominer library provides well-validated implementations of standard normalisation approaches.
Step 8 — Profile analysis and interpretation
Normalised morphological profiles can be used for compound clustering, mechanism of action deconvolution, toxicity prediction, disease phenotype characterisation, phenotypic reversion screening, and genetic interaction mapping.
The Science Behind pixlbio's Approach: Our Research
pixlbio's Cell Painting platform is built directly on the research of its scientific co-founders. Below are key publications from Jordi Carreras-Puigvert and Ola Spjuth that have shaped how we approach image-based phenomics in drug discovery.
Cell Painting: A Decade of Discovery — Nature Methods (2025)
Seal, Trapotsi, Spjuth, Singh, Carreras-Puigvert et al., Nature Methods, 2025 — DOI: 10.1038/s41592-024-02528-8
The definitive 10-year systematic review of Cell Painting, co-authored by Ola Spjuth and Jordi Carreras-Puigvert alongside Anne Carpenter and Shantanu Singh of the Broad Institute and AstraZeneca's imaging team. This paper reviewed 90 primary research articles spanning 2013–2023, synthesising advances in protocol development, feature extraction, machine learning integration, toxicity prediction, and mechanism of action deconvolution. It is the most comprehensive reference available for understanding where the field has come from and where it is going.
Combining Molecular and Cell Painting Data for MoA Prediction — AI in Life Sciences (2023)
Tian, Harrison, Sreenivasan, Carreras-Puigvert, Spjuth. Artificial Intelligence in the Life Sciences, 2023 — DOI: 10.1016/j.ailsci.2023.100060
This paper from our Uppsala group demonstrated that combining Cell Painting morphological profiles with molecular fingerprint data produces substantially better mechanism of action predictions than either data source alone. Using deep learning on both imaging and chemical structure data simultaneously, the study showed synergistic predictive power for 10 well-represented MoA classes — a key insight that informs how pixlbio builds predictive models today.
A Phenomics Approach for Antiviral Drug Discovery — BMC Biology (2021)
Rietdijk, Tampere, Spjuth, Carreras-Puigvert et al., BMC Biology, 2021 — DOI: 10.1186/s12915-021-01086-1
This paper, initiated by Ola Spjuth and Jordi Carreras-Puigvert, demonstrated that a modified Cell Painting protocol incorporating antibody-based viral detection could identify novel antiviral compounds and characterise virus-induced morphological signatures — demonstrating the adaptability of Cell Painting far beyond its original small molecule screening context. The approach achieved AUROC of 0.98 for distinguishing infected from non-infected cells.
CPSign: Conformal Prediction for Cheminformatics Modelling — Journal of Cheminformatics (2024)
Spjuth et al., Journal of Cheminformatics, 2024
Ola Spjuth's open-source conformal prediction framework CPSign — which underpins pixlbio's approach to uncertainty quantification in predictive modelling — provides calibrated, statistically valid confidence estimates for machine learning predictions. Unlike standard ML models that output point predictions, conformal prediction produces prediction intervals with guaranteed error rates. This matters enormously for drug safety applications, where knowing how confident a model is in a prediction is as important as the prediction itself. CPSign is available on GitHub and is in production use in multiple pharmaceutical organisations.
Why Cell Painting is Transforming Drug Discovery
Cell Painting's impact on drug discovery stems from several unique properties that distinguish it from conventional approaches.
It is unbiased by design. Unlike biochemical assays that measure a predefined target, Cell Painting captures the global morphological state of the cell — detecting unexpected biology, mechanisms, toxicities, and phenotypes that hypothesis-driven assays would miss entirely.
It is information-dense. A single experiment generates thousands of features per cell per condition — enabling statistical approaches that are simply not possible with conventional assays.
It scales to the library level. Once a Cell Painting pipeline is established, it can be applied to thousands of compounds with relatively modest incremental cost. Large-scale screens containing hundreds of thousands of compound profiles are now being used as reference maps against which new compounds can be positioned.
It integrates with modern AI. The numerical profiles produced by Cell Painting are ideal inputs for machine learning models. The combination of Cell Painting with deep learning — for both feature extraction and predictive modelling — is one of the most active areas of computational biology right now. Our own team's work on mechanism of action prediction and conformal prediction models is a direct expression of this integration.
Key Applications
Mechanism of Action Deconvolution
Compounds that share a mechanism of action produce similar morphological profiles. By comparing a new compound's profile against a reference library of known mechanisms, researchers can infer mechanism without prior hypotheses.
Early Toxicity Prediction
Morphological changes associated with cytotoxicity, hepatotoxicity, cardiotoxicity, and other adverse effects produce characteristic Cell Painting signatures — often detectable at sub-lethal concentrations, before conventional viability assays show any signal. At pixlbio, we apply this specifically to DILI prediction using our iPSC-derived hepatocyte platform. See our post on how we use iPSC hepatocytes and foundation models to decode DILI for more detail.
Disease Phenotype Characterisation and Phenotypic Reversion
Cell Painting can quantify the morphological difference between healthy and disease cell states — creating a phenotypic signature of disease that becomes a screening readout. This is directly relevant to our MASLD disease model work — where Cell Painting on iPSC-derived hepatocytes with PNPLA3 and TM6SF2 risk variants provides a human-relevant readout for phenotypic reversion screening.
Target Identification in Phenotypic Screens
When a compound produces a desired phenotypic effect, Cell Painting profiles from genetic perturbation screens can be mined to identify which gene knockouts produce a similar morphological profile — suggesting target engagement or pathway involvement.
Gene Function Annotation
Large-scale genetic perturbation screens using Cell Painting — such as the JUMP-CP dataset with 116,750+ compounds and 20,000+ genetic perturbations — are systematically revealing the morphological consequences of perturbing thousands of genes.
Cell Painting vs Other Profiling Approaches
Cell Painting occupies a unique position — high information content at high throughput and moderate cost. For liver biology specifically, combining Cell Painting with human-relevant iPSC-derived hepatocytes rather than cancer cell lines shifts the balance further — see our comparison of iPSC hepatocytes vs primary cells vs HepG2 for details on why cell model choice matters as much as the imaging platform.
Getting Started: Essential Resources
Foundational Papers
- Bray et al. (2016) — Cell Painting protocol. Nature Protocols. Read this first.
- Seal, Spjuth, Carreras-Puigvert et al. (2025) — Cell Painting: a decade of discovery. Nature Methods. The field's definitive 10-year review.
- Tian, Carreras-Puigvert, Spjuth et al. (2023) — Combining molecular and Cell Painting data for MoA prediction. AI in Life Sciences.
- Rietdijk, Spjuth, Carreras-Puigvert et al. (2021) — Phenomics approach for antiviral drug discovery. BMC Biology.
- Caicedo et al. (2017) — Data-analysis strategies for image-based cell profiling.
- Chandrasekaran et al. (2023) — CPJUMP1: three million images, 116,750 compounds.
Open Datasets
- JUMP-CP — The largest public Cell Painting dataset: 3M+ images, matched chemical and genetic perturbations
- RxRx datasets — Large-scale Cell Painting datasets from Recursion Pharmaceuticals
- Broad Bioimage Benchmark Collection (BBBC) — Curated reference datasets
- IDR (Image Data Resource) — Community repository for published biological image data
Software and Analysis Tools
- CellProfiler — Open-source standard for Cell Painting feature extraction
- pycytominer — Python library for morphological profile normalisation
- Cellpose — State-of-the-art deep learning cell segmentation
- DeepProfiler — Deep learning feature extraction for Cell Painting
- CPSign — Conformal prediction for cheminformatics modelling (Ola Spjuth, Uppsala)
Advanced Topics: The Frontier of Cell Painting
Foundation Models and Self-Supervised Learning
The field is rapidly moving beyond hand-crafted CellProfiler features towards deep learning foundation models — large neural networks pretrained on millions of Cell Painting images. Key developments include DINOv2 applied to Cell Painting, CellViT vision transformers, and DINO-based embeddings from the JUMP consortium pretrained on CPJUMP1. At pixlbio, our own foundation models trained on proprietary large-scale datasets are central to how we deliver predictive power to pharmaceutical customers. Foundation model embeddings consistently outperform CellProfiler features on MoA and toxicity prediction tasks — particularly for subtle phenotypic changes.
Cell Painting in 3D and Organoids
Standard protocols are optimised for 2D monolayer cultures. Extending to 3D systems — spheroids, organoids, tissue sections — introduces additional technical complexity but significantly greater biological relevance. This is an area our team is actively developing at pixlbio, particularly in the context of our iPSC-derived organoid models.
Integrating Cell Painting with Other Omics
Multi-modal profiling — combining Cell Painting with transcriptomics, proteomics, or metabolomics — captures complementary biological information. The JUMP-CP consortium's matched perturbation dataset is an early example at scale. Our work combining molecular fingerprints with morphological profiles for MoA prediction is directly in this tradition.
Cell Painting for iPSC Disease Models and Precision Medicine
As iPSC technology enables patient-derived cell models from individuals with specific genetic variants, Cell Painting can characterise morphological consequences of disease mutations and screen for compounds that reverse disease phenotypes in genotype-specific models. This is the foundation of pixlbio's disease model platform — combining CRISPR-engineered iPSC hepatocytes carrying MASLD risk variants, A1ATD mutations, or PFIC2 genotypes with Cell Painting readouts to enable human-relevant phenotypic drug discovery.
Common Challenges and How to Address Them
Batch effects. Systematic differences between experimental runs are the most common technical challenge at scale. Mitigation: consistent plate layouts with well-distributed controls, strict standardisation of reagent lots and imaging conditions, and computational correction using spherize normalisation or ComBat.
Cell type optimisation. The standard protocol was developed primarily in U2OS cancer cells. Applying it to primary cells, iPSC-derived models, or organoids requires re-optimisation — particularly for staining concentrations, fixation conditions, and segmentation.
Data storage and compute. A 384-well experiment generates 50–200 GB of raw image data. Cloud-based solutions are increasingly standard for large-scale campaigns.
Interpretability. Cell Painting profiles are high-dimensional numerical vectors — not easy to interpret directly. Models with built-in uncertainty quantification, such as the conformal prediction approach Ola Spjuth's group has developed, help address this directly.
How pixlbio Approaches Cell Painting
We built pixlbio's pixCellPaint service because we saw a consistent gap in the field: researchers understood the value of Cell Painting but lacked the infrastructure, expertise, or cell models to run it reliably at scale.
Our approach is deliberately model-agnostic and perturbation-agnostic. We work with any cell type relevant to your biology — our own iPSC-derived models, customer-supplied cells, primary cells, or established cell lines — and any perturbation type or biological question.
What we bring is the infrastructure, expertise, and AI analytics to turn raw Cell Painting data into actionable biological insights. Our team includes scientists who have contributed directly to the academic literature on Cell Painting methodology — including the Nature Methods 10-year review — and who apply that knowledge to every study we run.
The combination of human-relevant cell models with Cell Painting phenomics and AI-driven analysis is, in our view, the most powerful available approach to preclinical drug discovery today.
The pixlbio Cell Painting platform was designed and built by scientific co-founders Jordi Carreras-Puigvert and Ola Spjuth. Jordi is CSO & Co-Founder of pixlbio and Associate Professor at Uppsala University, where his research focuses on image-based phenomics, high-content screening, and AI for drug discovery. He is co-author of the 2025 Nature Methods Cell Painting review alongside Anne Carpenter and Shantanu Singh of the Broad Institute. Ola is CAIO & Co-Founder of pixlbio and Professor of Pharmaceutical Bioinformatics at Uppsala University, where he leads the Pharmb.io research group. With 5,600+ citations, he is one of the foremost researchers in AI for drug discovery and conformal prediction, and creator of CPSign. Together they have co-authored landmark papers on Cell Painting, mechanism of action prediction, and AI-driven phenomics — including the 2025 Nature Methods review, the 2023 MoA prediction paper, and the 2021 antiviral phenomics paper.
Conclusion
- Seal, Trapotsi, Spjuth, Singh, Carreras-Puigvert et al. (2025) Cell Painting: a decade of discovery and innovation in cellular imaging. Nature Methods, 22(2):254–268. DOI: 10.1038/s41592-024-02528-8
- Tian, Harrison, Sreenivasan, Carreras-Puigvert, Spjuth (2023) Combining molecular and cell painting image data for mechanism of action prediction. AI in Life Sciences. DOI: 10.1016/j.ailsci.2023.100060
- Rietdijk, Tampere, Spjuth, Carreras-Puigvert et al. (2021) A phenomics approach for antiviral drug discovery. BMC Biology. DOI: 10.1186/s12915-021-01086-1
- CPSign — Spjuth et al. (2024) Conformal prediction for cheminformatics modeling. Journal of Cheminformatics.
- Bray et al. (2016) Cell Painting, a high-content image-based assay for morphological profiling. Nature Protocols.
- Nikolaou et al. (2024) Development of optimised human iPSC-derived hepatocytes. bioRxiv.
- Wawer et al. (2014) — Toward performance-diverse small-molecule libraries for cell-based phenotypic screening using multiplexed high-dimensional profiling. PNAS, 111(30):10911–10916.